Just a quick note at the start of this post. This was the initial version of MNCA. For the latest code and examples make sure you check out this post and this post too.
This one comes from the code here from u/slackermanz. He didn’t have a name for them at the time so I coined the term “Multiple Neighborhood Cellular Automata“.
For this CA there are 4 different large size neighborhoods used each step.
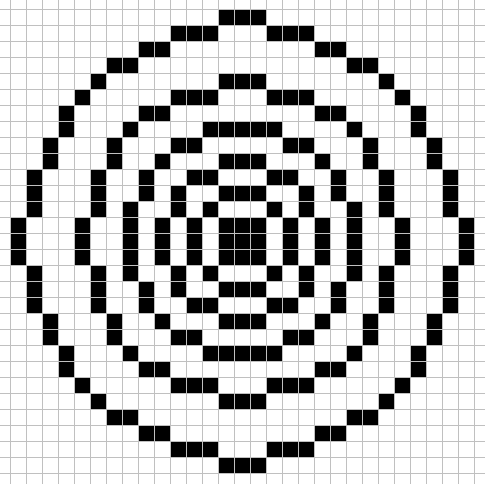
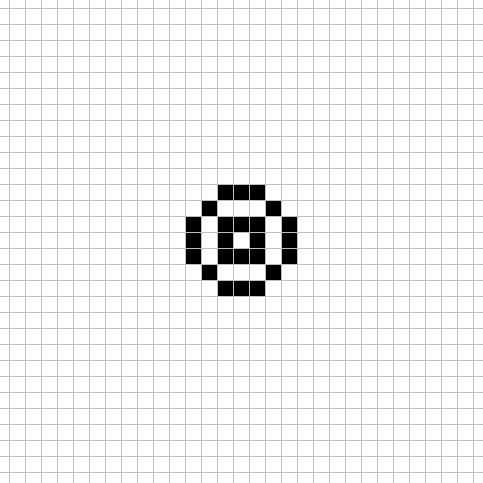
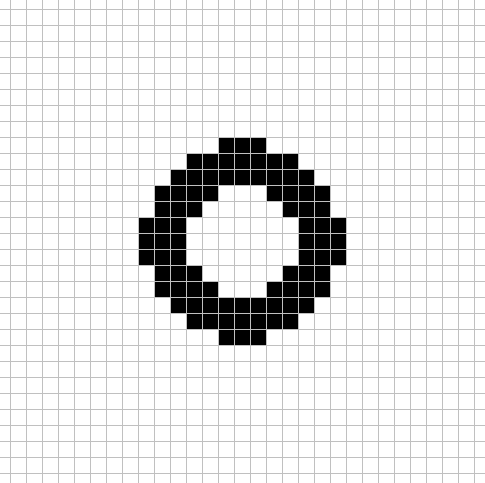
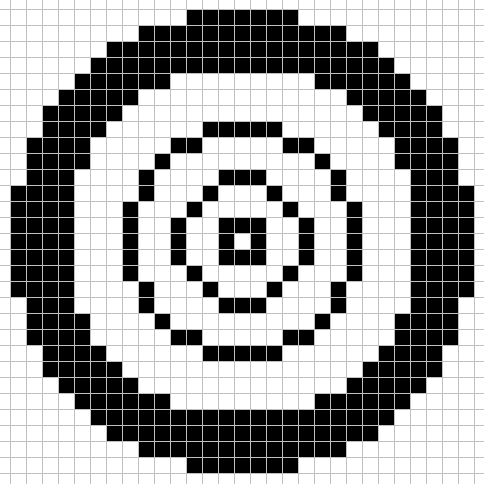




Each cell uses the above neighborhood patterns to tally the live cells into 4 sum values. sum_0 is the live cell count in neighborhood 1, sum_1 is the live cell count in neighborhood 2, sum_2 is the live cell count in neighborhood 3, and sum_3 is the live cell count in neighborhood 4.
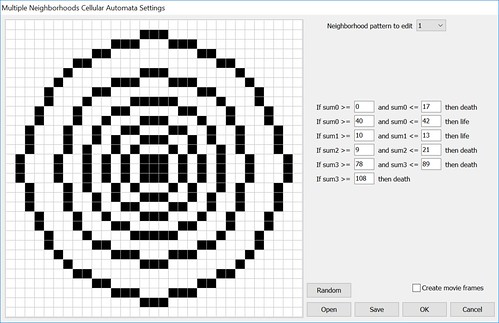
The sums are used to determine life or death for the cells by using the following formulas.
If sum_0 is between 0 and 17 then the cell dies.
If sum_0 is between 40 and 42 then the cell lives.
If sum_1 is between 10 and 13 then the cell lives.
If sum_2 is between 9 and 21 then the cell dies.
If sum_3 is between 78 and 89 then the cell dies.
If sum_3 is greater than 108 then the cell dies.
Put all that into an options form to allow easy config changes.
Here is a quick sample movie with a few of the interesting rules I found so far.
Multiple Neighborhoods Cellular Automata are now included in the latest version of Visions of Chaos.
See here and here for more explorations into the world of Multiple Neighborhood Cellular Automata.
Jason.
Hello, With MNCA, there seems to be a Bug, the software hangs at 37 ”.
37 seconds? Using the default startup config? MNCA does use the GPU for calculations and rendering so try updating your graphics card drivers. That may help.
Thank you, I’m going to look at this
Pingback: Understanding Multiple Neighborhood Cellular Automata - Slackermanz
I haven’t tested if it’s faster, and I don’t know if you’re doing this already, but I implemented this using convolutions with the neighbourhoods to find the number of neighbours. In pytho, scipy uses fft to do convolutions for larger images so it should be faster than just checking each cell. I also use a single convolution kernel with a different channel for each neighbourhood so I only need to convolve once and get a matrix with the number of all neighbouring cells for each different neighbourhood.